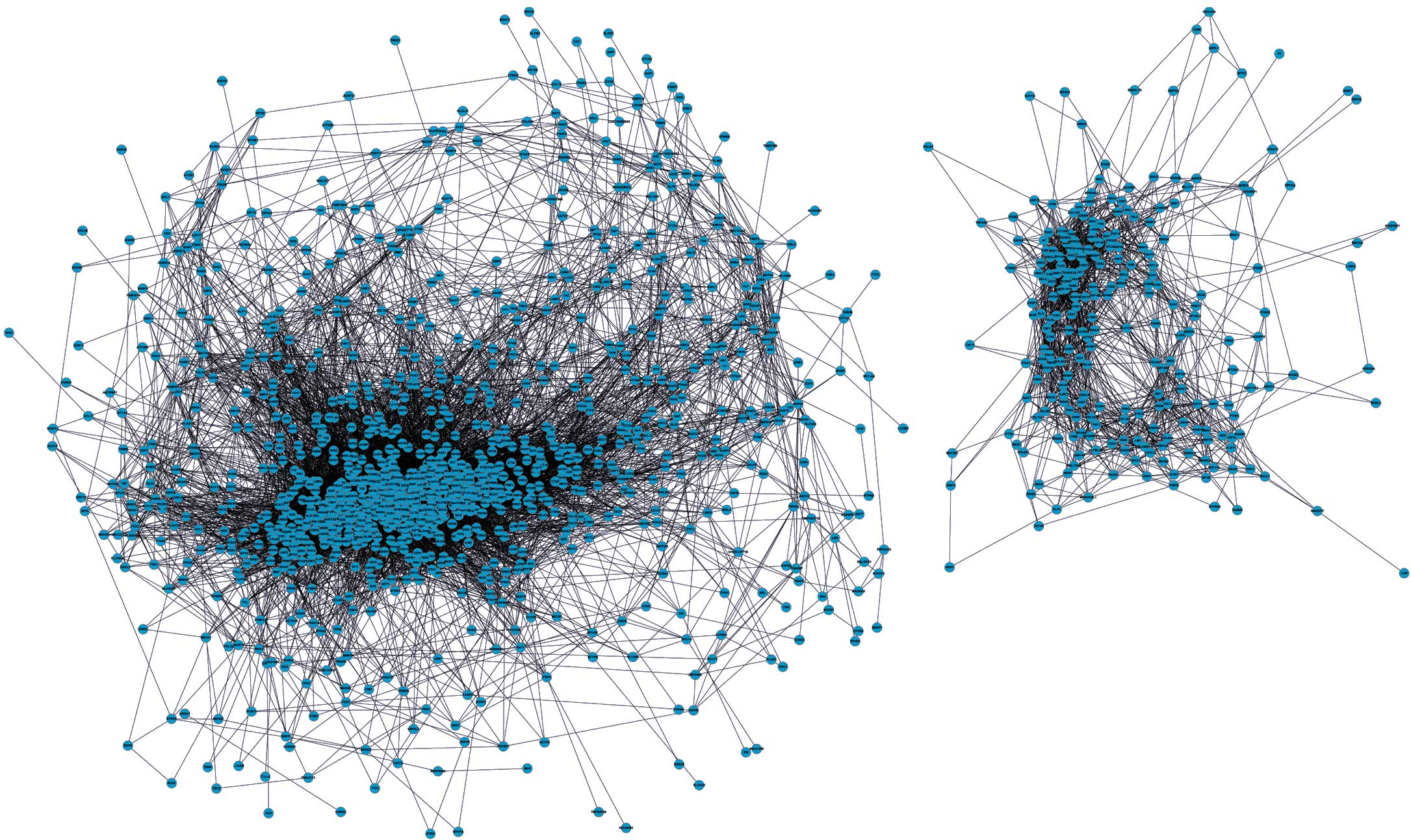
Expression of CTNNA2 was decreased in MYOG knock-down cells. To confirm this finding, in vitro studies revealed increased expression of CTNNA2 in MSCs on day 12 compared to day 10. Four functional clusters (modules or communities) were identified in the network and the functional enrichment analysis revealed that genes included in these clusters significantly contribute to skeletal muscle development. By constructing an interaction network of differentially expressed genes (DEGs) using GeneMANIA, cadherin-associated protein (CTNNA2) was identified as the main hub gene in the network with highest node degree. Using an RNA-Seq approach, analysis of MYOG knock-down muscle satellite cells (MSCs) have shown that genes associated with cell cycle and division, DNA replication, and phosphate metabolism are differentially expressed. Despite a plethora of information available regarding MYOG, the mechanism by which MYOG regulates muscle cell differentiation has not yet been identified. It is evident that MYOG function cannot be compensated by any other myogenic regulatory factors (MRFs). Subsequent to a decline in Pax7, induction in the expression of MYOG is a hallmark of myoblasts that have entered the differentiation phase following cell cycle withdrawal. Muscle, a multinucleate syncytium formed by the fusion of mononuclear myoblasts, arises from quiescent progenitors (satellite cells) via activation of muscle-specific transcription factors (MyoD, Myf5, myogenin: MYOG, and MRF4).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
